JupyterLab
Next-generation Web interface

JupyterLab is an advanced, extensible, and feature-rich application for creating and editing computational notebooks, integrating code, natural-language explanations, data, and visualizations.
It is part of Project Jupyter, an initiative that provides standards and tools for interactive computing and reproducible science.
Although accessed through the browser, it is not just a “web interface”, but a complete working environment with multiple panels, extensions, support for different languages, and integration with data and science ecosystems.
Officially released in 2018, JupyterLab was funded by organizations such as the Gordon and Betty Moore Foundation and the Alfred P. Sloan Foundation, aiming to expand Jupyter tooling and deliver a more complete interactive development experience.
While a “sibling” to other notebook authoring apps under the Project Jupyter umbrella (e.g., Jupyter Notebook and Jupyter Desktop), it provides a more advanced, richer, and customizable experience, standing out for its extensibility, modularity, and broad adoption in scientific and educational communities.
What it does
- Notebook authoring: combines executable code cells, explanatory text (Markdown/LaTeX), visualizations, and interactive results.
- Unified environment: open notebooks, terminals, text files, JSON, CSV, and Markdown within the same window.
- Interactive execution: connects to kernels for multiple languages (Python, R, Julia, Scala, etc.).
- Data exploration: supports advanced visualization libraries (Matplotlib, Plotly, Bokeh, Vega, 3D).
- Sharing & reproducibility: notebooks are portable and versionable documents.
- Extensibility: add dashboards, special format renderers, Git integration, data connectors, and more.
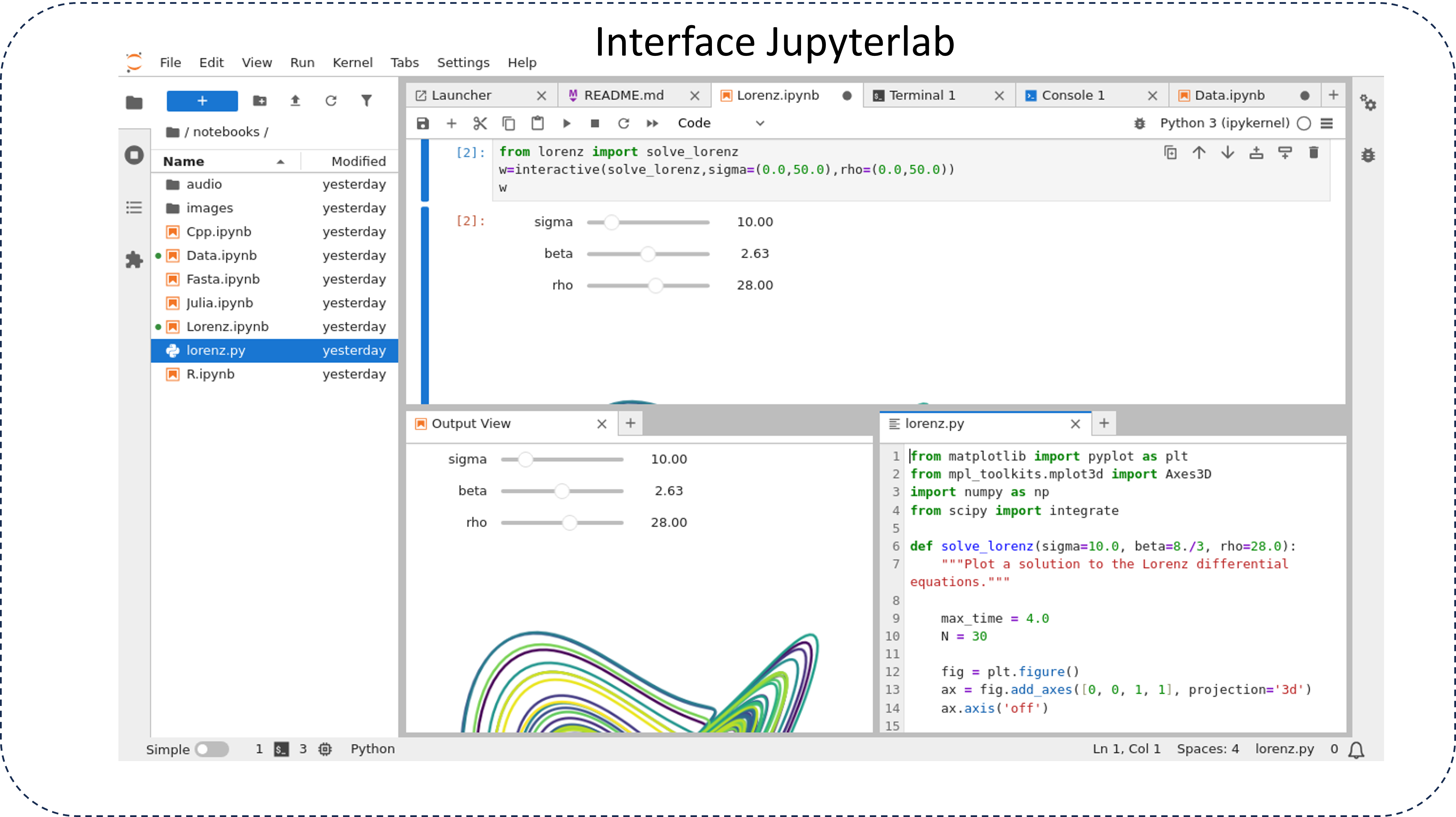
JupyterLab features:
- Modular interface: multiple tabs and draggable panels, customizable layouts.
- Command palette: central shortcut hub, similar to modern IDEs.
- Kernel & session management: view, restart, or stop active kernels.
- File manager: navigate and manipulate server-side directories.
- Embedded terminals: integrated shell access.
- Extension integration: from new data formats to custom panels.
- Backward compatibility: older Jupyter notebooks remain functional.
JupyterLab architecture
JupyterLab is built on a layered architecture.
- Frontend (JupyterLab UI): implemented in TypeScript, based on modular widgets, runs in the browser, and manages user interaction.
- Jupyter Server: backend responsible for:
- Managing files and directories.
- Serving the web application.
- Brokering WebSockets/HTTP communication between the UI and kernels.
- Kernels: independent processes for executing code in different languages. The most popular is the IPython Kernel, with many others (R, Julia, Scala, Java, Rust).
- Messaging protocols: communication between UI and kernels follows the Jupyter Messaging Protocol, based on JSON and ZeroMQ, standardizing requests, responses, and events.
Main components:
- Notebook: interactive document with code and markdown cells.
- Code console: advanced REPL connected to a kernel, useful for quick tests.
- Text editor: with syntax highlighting, useful for scripts, JSON, configs.
- Terminal: Unix shell inside the browser.
- File Browser: manages files, directories, and drag-and-drop.
- Launcher: start screen to open new notebooks, consoles, or files.
- Extensions: third-party or official modules that expand UI and functionality.
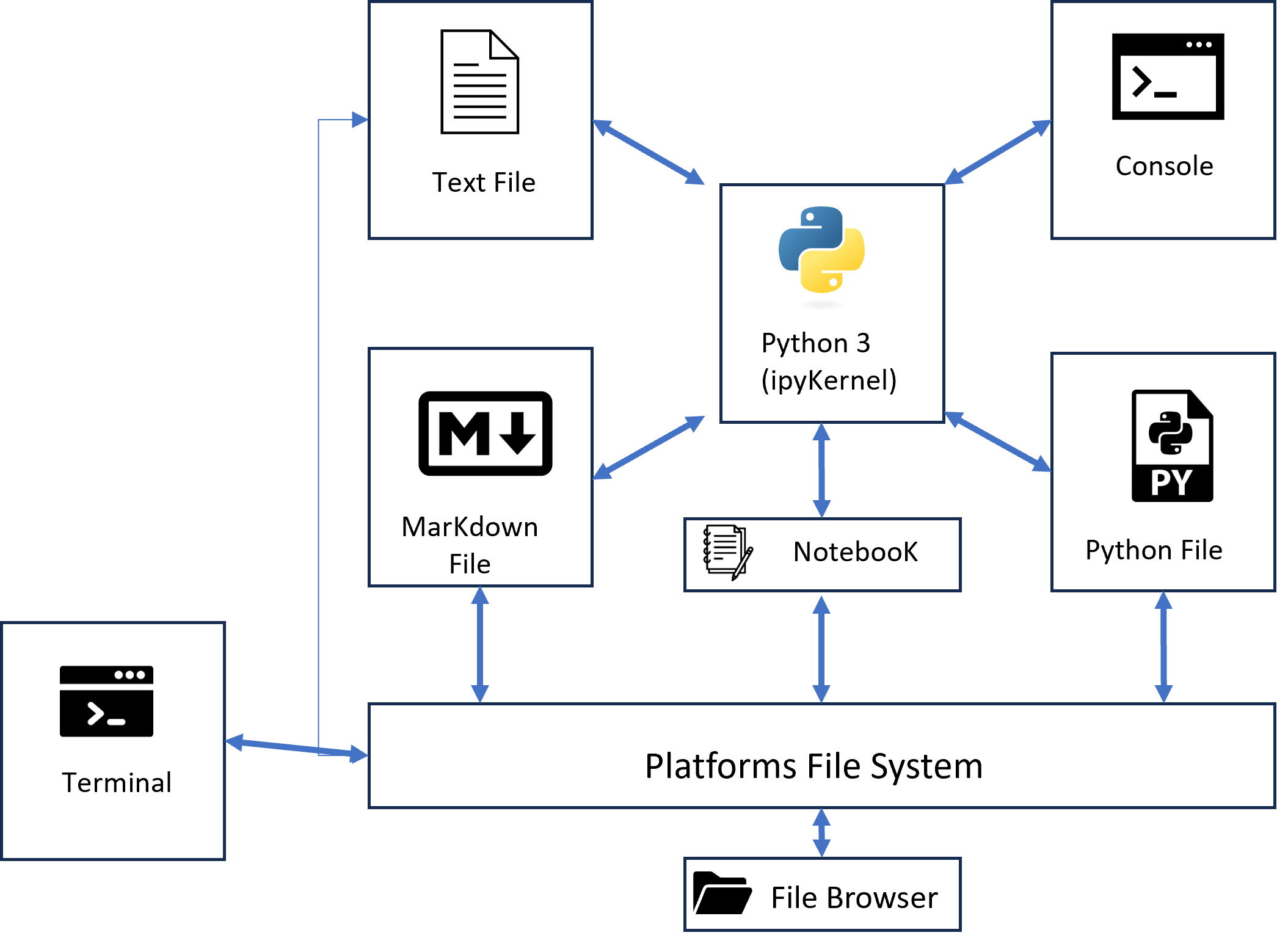
JupyterLab capabilities
- Flexible layouts: arrange notebooks, consoles, and files side by side.
- Advanced visualizations: interactive charts, 3D models, dashboards.
- Multi-language support: kernels for dozens of languages.
- Real-time collaboration: with JupyterLab RTC (experimental).
- Export: convert notebooks to PDF, HTML, slides (via nbconvert).
- Git and CI/CD integration: extensions enable built-in versioning.
Good practices when using JupyterLab
- Manage environments: keep each project’s dependencies in an isolated, reproducible environment using tools suited to your ecosystem (environment managers, packagers, or containers).
- Document notebooks: combine code with clear explanations, titles, and descriptions.
- Avoid heavy outputs: clear outputs before committing to keep repositories small.
- Version control: integrate with Git; notebooks render natively on GitHub.
- Security: do not run notebooks from unknown sources without review.
- Modular structure: use notebooks to prototype; when stable, move functions to Python modules.
- Export & reporting: use conversion tools to publish notebooks as PDF, slides, or HTML (e.g., nbconvert, Quarto, Jupyter Book).
- Be cautious with extensions: install only what you need and keep them updated.
When to use JupyterLab
-
Rapid prototyping of code and ideas Test snippets interactively with immediate feedback—ideal for data exploration, ad-hoc analysis, and algorithm trials. Keep code + explanation + result in one place.
-
Data analysis and exploration Great for light ETL, exploratory charts, and quick statistics. Integrates with Pandas, NumPy, Matplotlib, Plotly, Seaborn, Bokeh. Widgets and parameterized notebooks enrich exploration beyond static scripts.
-
Education, training, and technical documentation Perfect for classes and tutorials: read theory, run code, and see results in one flow. Widely used in MOOCs, bootcamps, and data-science e-books. Enables reproducible materials.
-
Reproducible science and academic research Document analytical steps (code + data + explanations). Export to papers (PDF, LaTeX, HTML). Reproduce experiments, share pipelines, and embed visualizations.
-
Collaboration in data teams Shared environments (on-prem or cloud) can expose JupyterLab as a notebook hub. With JupyterHub or collaboration extensions, multiple people access the same server—reducing “works on my machine” issues.
-
Projects with multiple files and languages Unlike classic Jupyter Notebook, JupyterLab opens multiple files side by side (notebook + CSV + Markdown + terminal). Useful when editing helper scripts, config JSON, or Markdown without leaving the environment.
-
When extensibility is required Customize the interface or integrate external tools (Git, MLflow, dashboards, specific renderers). The extension ecosystem makes JupyterLab a lightweight, notebook-first IDE.
When not to use JupyterLab (practical limitations)
- Large/complex software development: for robust projects with many modules and tests, IDEs like PyCharm or VS Code offer better support.
- Very long-running jobs in notebooks: if code runs for hours/days, prefer versioned scripts or pipelines (Airflow, Prefect, Luigi).
- Production/Deploy: JupyterLab is for prototyping and analysis, not continuous production execution.
Project details
In essence, while Python is the dominant language for data science and computation within JupyterLab, TypeScript (with React and Lumino, formerly PhosphorJS) plays a crucial role in building the platform’s frontend and in extending its capabilities.
The backend is developed in Python, serving files and the web UI, managing sessions and authentication, and brokering communication with kernels via the Jupyter protocol.
Regarding kernels, the IPython kernel is the most widely used (also written in Python) and is extensible to many languages (R, Julia, Scala, Java, Rust, etc.).
TDP Kubernetes
This component is also available in the TDP Kubernetes edition (from version 3.0.0) with version 5.3.0, deployed via Helm Chart tdp-jupyter.
For configuration details, see the JupyterHub documentation in TDP Kubernetes.
Sources: